Multicollinearity
Multicollinearity occurs when predictor variables are highly correlated with each other. It makes individual variable effects difficult to estimate reliably: coefficients become unstable, standard errors inflate, and interpretation becomes ambiguous.
It is important to distinguish between two situations:
- Inference (which variables matter, what drives the pattern): multicollinearity is a serious problem
- Prediction (how well can I predict the outcome): multicollinearity is much less of a concern
Common examples in biology include body weight and body length, genes in the same pathway, temperature and humidity, or multiple measures of the same trait such as leaf length and leaf area.
Detection
Correlation matrix
A natural first step after computing correlations. Pairwise values above roughly |r| = 0.8 are worth investigating:
library(corrplot)
corrplot(cor(mtcars),
method = "number",
type = "upper",
tl.col = "black",
tl.srt = 45)
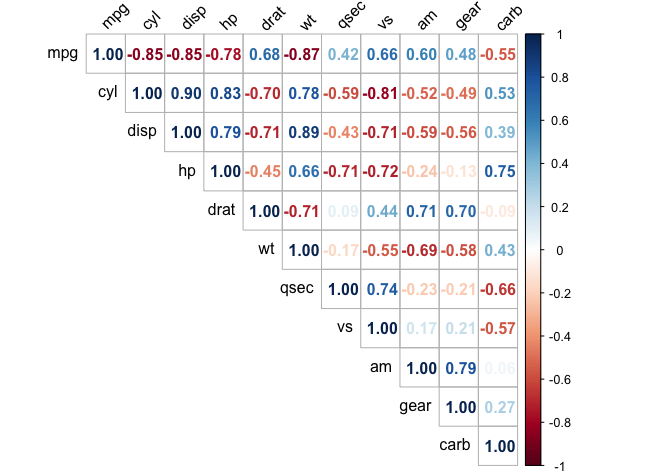
Variance Inflation Factor (VIF)
VIF is the standard diagnostic for regression models. It quantifies how much the variance of a coefficient is inflated due to collinearity with other predictors:
library(car)
model <- lm(mpg ~ cyl + disp + hp + wt, data = mtcars)
round(car::vif(model), 2)
#> cyl disp hp wt
#> 6.74 10.37 3.41 4.85
Both cyl and disp have VIF values well above 10, which indicates severe collinearity. Their individual effects on mpg cannot be reliably separated from one another. In contrast, hp and wt fall within an acceptable range and can remain in the model as they are.
| VIF | Interpretation |
|---|---|
| 1 | No collinearity |
| 2 to 5 | Moderate, usually acceptable |
| 5 to 10 | High, consider action |
| > 10 | Severe, action needed |
Solutions
Which solution fits depends on your goal:
graph TD
A[Multicollinearity detected] --> B{Primary goal?}
B -->|Interpretation| C{How many variables?}
B -->|Prediction| D[PCA or Ridge regression]
C -->|Few| E[Remove or combine variables]
C -->|Many| F[Lasso regression]Remove one variable from a correlated pair. Keep the one with the clearer biological interpretation. In the mtcars example, cyl and disp both measure engine size, so keeping disp alone brings the VIF down to acceptable levels.
model2 <- lm(mpg ~ disp + hp + wt, data = mtcars)
round(car::vif(model2), 2)
#> disp hp wt
#> 7.32 2.74 4.84
Combine variables into a single composite. This works best when two predictors measure the same underlying quantity. Multiplying leaf length and leaf width gives an approximation of leaf area, replacing two correlated predictors with one ecologically meaningful variable. The composite should always have a clear interpretation.
PCA replaces a set of correlated variables with uncorrelated principal components. This is the most principled solution when many variables are involved, but the components are linear combinations of the originals and can be harder to interpret biologically. In the mtcars example, the first component typically captures a general engine size gradient explaining over 80% of the variance among the four predictors:
pca <- prcomp(mtcars[, c("cyl", "disp", "hp", "wt")], scale. = TRUE)
summary(pca) # PC1 usually explains > 80% here
model <- lm(mpg ~ pca$x[, 1] + pca$x[, 2], data = mtcars)
# Coefficients now refer to components, not the original variables
Regularised regression stabilises estimates by penalising large coefficients without removing variables beforehand. Ridge shrinks all coefficients toward zero but retains every predictor, which makes it useful for prediction. Lasso sets some coefficients exactly to zero, effectively selecting a subset of variables and making results more interpretable:
library(glmnet)
X <- as.matrix(mtcars[, c("cyl", "disp", "hp", "wt")])
y <- mtcars$mpg
cv_ridge <- cv.glmnet(X, y, alpha = 0) # Ridge: keeps all variables
cv_lasso <- cv.glmnet(X, y, alpha = 1) # Lasso: excludes some variables
coef(cv_lasso, s = "lambda.min")
# Coefficients set to exactly zero are excluded from the model
| Strategy | Variables kept | Interpretability | Best for |
|---|---|---|---|
| Remove | Fewer | High | Few correlated pairs |
| Combine | Fewer | Medium | Biologically meaningful composites |
| PCA | Components | Lower | Many correlated variables |
| Ridge | All | Medium | Prediction |
| Lasso | Some | High | Inference with many predictors |
Exercise
Using the iris dataset, build a model predicting Sepal.Length from the other three numeric variables. Detect multicollinearity and apply an appropriate solution.
Solution
library(car)
model <- lm(Sepal.Length ~ Sepal.Width + Petal.Length + Petal.Width,
data = iris)
round(car::vif(model), 2)
#> Sepal.Width Petal.Length Petal.Width
#> 1.27 15.10 14.23
# Petal.Length and Petal.Width are severely collinear
# Option 1: remove the less interpretable variable
model2 <- lm(Sepal.Length ~ Sepal.Width + Petal.Length, data = iris)
vif(model2) # Both now near 1.3
# Option 2: replace both with a PC
pca <- prcomp(iris[, 3:4], scale. = TRUE)
model3 <- lm(Sepal.Length ~ iris$Sepal.Width + pca$x[, 1], data = iris)